Promotion Details (valid for Americas and Europe customers NOW through July 31)
A. The 899 USD/sample package - 50X human exome sequencing
- Agilent SureSelect 50/51M Capture kit
- 100 bp paired-end sequencing on HiSeq 2000
- 5 Gb high quality* sequencing data
- 50X average coverage for target regions guaranteed
- ~85% targeted bases with sequencing depth >= 10X for qualified samples
- SNP & Indel calling and annotation included
B. The 1299 USD/sample package - 100X human exome sequencing 
- Agilent SureSelect 50/51M Capture kit
- 100 bp paired-end sequencing on HiSeq 2000
- 10 Gb high quality* sequencing data
- 100X average coverage for target regions guaranteed
- ~90% targeted bases with sequencing depth >= 10X for qualified samples
- SNP & Indel calling and annotation included
*High-quality data are generated from raw data with adaptor and low quality sequences removed.
Lab location: BGI Hong Kong, the world’s highest sequencing throughput at one site
Turnaround time: BGI can complete the library construction, exome capture, sequencing, and analysis within 6-8 weeks for up to 400 samples
Minimum order: 20 samples. The first batch of samples must be received by BGI within 2 months after contract signing.
Sample requirements: 1 μg (3 μg gDNA recommended)
Offer valid for human DNA samples for Americas and Europe customers only NOW through July 31.
Why choose BGI?
- High quality data
- Highly experienced: More than 38,000 whole human exomes sequenced by BGI to date
- Rapid turnaround
- Affordable pricing
- Strong analytical capabilities: 1,000 bioinformaticians (Advanced analysis available for Mendelian disorders, complex diseases, cancer, and population analysis)
Why choose 50X and 100X?
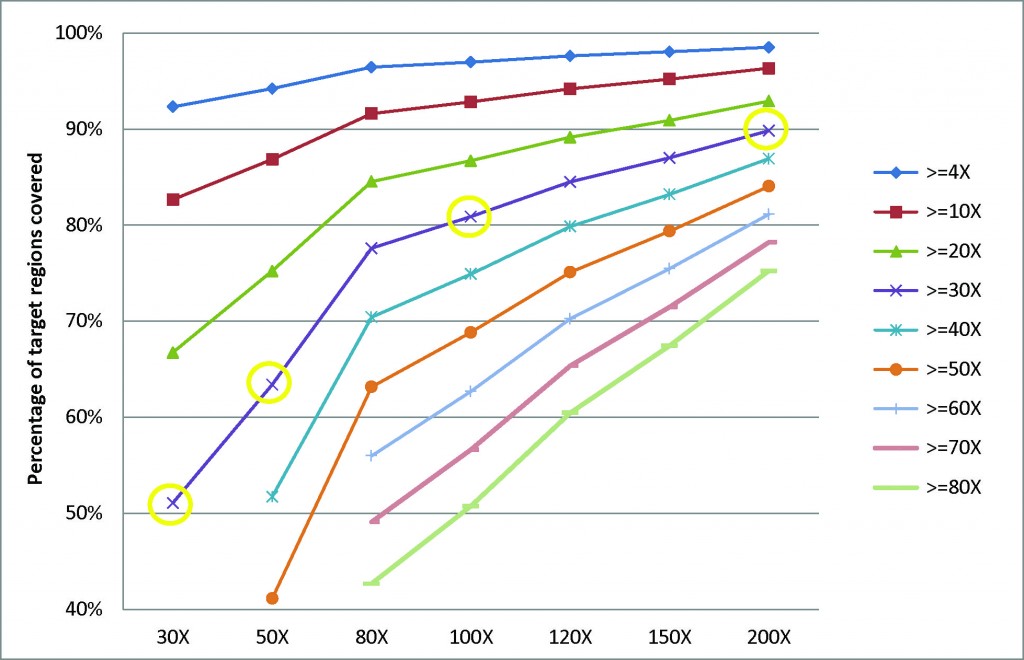
- Researchers can get better coverage and discover more variants as we increase the sequencing depth (Fig 1); therefore, gaining more insights into the genetic basis of various kinds of cancers and complex diseases.
Fig.1 Correlation between the percentage of target regions covered and the sequencing depth in human exome sequencing. Take >=30X series (the purple line) for example: when the sequencing depth is 30X, only half of the target regions (51%) are covered at above 30X. As we increase the sequencing depth, a much higher percentage (63% for 50X, 81% for 100X, and 90% for 200X) of the target regions is covered at above 30X. The data used in the analysis for each coverage level listed on the X axis are generated by randomly extracting, from a 200X exome sequencing result, the amount of data equivalent to the corresponding coverage level.
Our Publications (selected):
- Guo, Guangwu, et al. Frequent mutations of genes encoding ubiquitin-mediated proteolysis pathway components in clear cell renal cell carcinoma. Nature Genetics. 44, 17–19 (2012).
- Chiang, Pei-Wen, et al. Exome sequencing identifies NMNAT1 mutations as a cause of Leber congenital amaurosis. Nature Genetics. 44, 972–974 (2012).
- Xu, Xun, et al. Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumor. Cell. 148 ( 5), 886-895 (2012).
- McVean, Gil A., et al. An integrated map of genetic variation from 1,092 human genomes. Nature. 491, 56–65 (2012).
- Yi, Xin, et al. Sequencing of 50 human exomes eeveals adaptation to high altitude. Science. 329 (5987), 75-78 (2010).