- Genomics
- Transcriptomics
- Epigenomics
- Meta-omics
- Proteomics
- Single-Cell Sequencing
- Immune Repertoire Sequencing
- FFPE Samples

Protein Identification
Cases
Technical Information
Contact Us / Wish List
 Protein identification can be performed on a variety of samples, including protein gel spots, protein gel bands, IP samples, etc. Proteins are analyzed either by MALDI-TOF/TOF or by nano-LC-ESI-MS/MS and then identified using peptide masses or MS/MS fragmentation spectra. We provide your protein identification results in an electronic report that also includes bioinformatics analysis results, such as GO and pathway analysis.
Protein identification can be performed on a variety of samples, including protein gel spots, protein gel bands, IP samples, etc. Proteins are analyzed either by MALDI-TOF/TOF or by nano-LC-ESI-MS/MS and then identified using peptide masses or MS/MS fragmentation spectra. We provide your protein identification results in an electronic report that also includes bioinformatics analysis results, such as GO and pathway analysis.
Benefits:
- High resolution and accuracy: With LC-MS/MS, the accurate identification results can be provided based on resolving power up to 100,000 FWHM and 1 ppm mass accuracy.
- Ultra-fast scanning speed: UltrafleXtremeTM MALDI-TOF/TOF MS with 1 kHz smartbeam IITM laser technology can provide fast spectra collection in both TOF and TOF/ TOF modes.
- High sensitivity: Able to detect proteins at even nanogram levels.
Services:
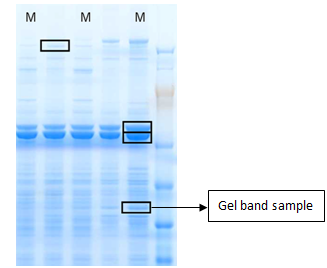
Gel Band/Spot Identification of Proteins
Identification of proteins from visible bands or spots excised from Coomassie or silver-stained gels using mass-spectrometry is generally very successful.
- Identification of unknown proteins in 1D or 2D gels
- Verification of protein expression products
- Detection of binding partners in immunoprecipitation experiments
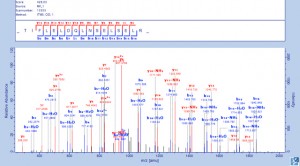
Gel Band Identification of Proteins by LC-MS/MS
Tarver MR, et al. J Insect Sci. 2012
Gel Spot Identification of Proteins by MALDI-TOF/TOF
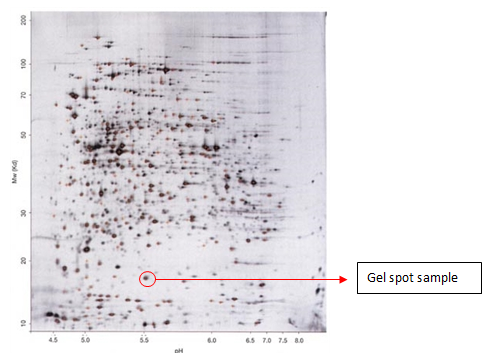

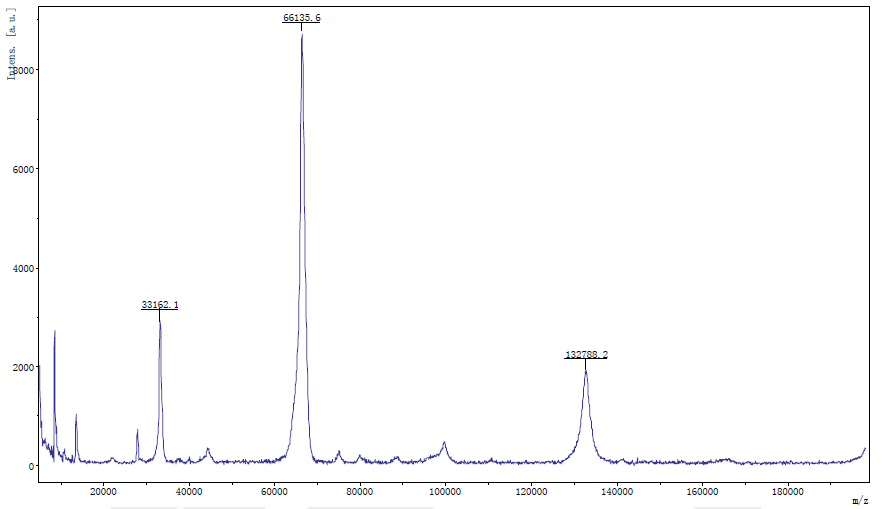
Protein Molecular Weight Detection
Determination of molecular weight of single or mixed protein by MALDI-TOF
Proteomics Study of Rice Embryogenesis: Discovery of The Embryogenesis-dependent Globulins. Electrophoresis. 33(7):1129-38 (2012).
The plant embryo is the germination center of the seed. Generally, the complex processes of embryogenesis result from the action of a coordinated network of genes. Thus, a large-scale survey of changes in protein abundance during embryogenesis is an effective approach to study the molecular events of embryogenesis. In this study, two-dimensional gel electrophoresis (2DE) was applied to separate rice embryo proteins collected during the three phases of embryogenesis: 6 days after pollination (DAP), 12 DAP, and 18 DAP. We then employed matrix-assisted laser desorption-ionization time of flight/time of flight mass spectrometry (MALDI TOF/TOF MS) to identify the phase-dependent differential 2DE spots. A total of 66 spots were discovered to be regulated during embryogenesis, and of these spots, 53 spots were identified. These proteins were further categorized into several functional classes, including storage, embryo development, stress response, glycolysis, and protein metabolism.
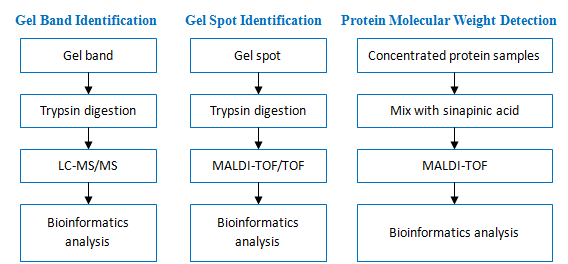
Workflow:
Bioinformatics:
Standard bioinformatics analysis
- Statistical analysis and quality control
- Protein identification
- Protein GO analysis (only for gel band identification)
- Protein COG analysis (only for gel band identification)
- Protein pathway analysis (only for gel band identification)
Sample Requirements:
- Gel band: the amount of proteins ≥30µg, concentration ≥1µg/µl. The gel band needs to be visible with coomassie blue or silver staining.
- Gel spot: the amount of proteins ≥50pmol. Salt content: volatile inorganic salt < 20mM, stable inorganic salt < 5mM.
- Protein molecular weight detection:
- Sample type: purified protein solution or powder (the purity of the sample should be > 90%)
- Total amount of protein: ≥50ug
- Protein concentration: ≥1ug/ul
- If silver staining method is used, please be sure not to fix the gel with glutaraldehyde.