
Microbial
 Microbes make up the majority of the earth’s biomass and have evolved for over 3.8 billion years. Knowledge of the enormous range of microbial capacities can have far-reaching benefits in medical, industrial, agricultural and environmental applications. By employing a combination of various cutting-edge sequencing technologies, such as Solexa, 454, MiSeq, and Ion poton, we have completed genome research on archaea, fungi, bacteria, protozoa, viruses as well as micro-ecological systems with more than 21,000 samples. Moreover, at least 80 single bacteria-related articles and 10 metagenome-related articles were published. These microbial genomes have been applied in a wide range of applications, for example, in comparative genomics and biomarkers development (Figure 1). BGI is also highly experienced in applying transcriptomics and proteomics to microbiology studies, and has generated a large amount of data to facilitate microbiology applications in medicine, agriculture and industry.
Microbes make up the majority of the earth’s biomass and have evolved for over 3.8 billion years. Knowledge of the enormous range of microbial capacities can have far-reaching benefits in medical, industrial, agricultural and environmental applications. By employing a combination of various cutting-edge sequencing technologies, such as Solexa, 454, MiSeq, and Ion poton, we have completed genome research on archaea, fungi, bacteria, protozoa, viruses as well as micro-ecological systems with more than 21,000 samples. Moreover, at least 80 single bacteria-related articles and 10 metagenome-related articles were published. These microbial genomes have been applied in a wide range of applications, for example, in comparative genomics and biomarkers development (Figure 1). BGI is also highly experienced in applying transcriptomics and proteomics to microbiology studies, and has generated a large amount of data to facilitate microbiology applications in medicine, agriculture and industry.
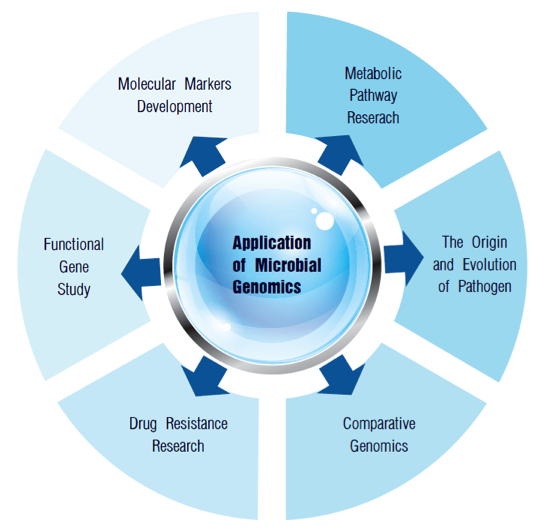
Figure 1. Applications of microbiology genome sequencing
Medical:
Increasingly, the value of the relationship between microbe populations living in the human body and human health is being recognized. Research has shown diverse diseases, such as obesity, enteritis, diabetes, and colon cancer, are impacted by microbe populations resident in the body. NGS opens the realm of incorporating routine whole-genome analyses in the study of pathogenic bacteria, viruses and parasites for infectious disease research studies. For our customers, we have both the expertise and leading technology platforms to provide reliable pathogen genome sequencing and bioinformatics analysis. We have performed sequencing studies of many pathogenic microbes from single strains to population levels. In 2010, we established the human gut microbial gene catalogue, the first human microbial gene catalogue derived by metagenomics sequencing. One year later, we sequenced genome of deadly E.coli within 3 days upon receipt of the bacteria samples during the E.coli outbreak in Germany, and thus help control the spread of the infection (Figure 2 and 3, paper). In 2013, we identified 85 genes and intergenic regions associated with drug resistance through genome sequencing of 161 Mycobacterium tuberculosis isolates from China (Figure 4, paper).
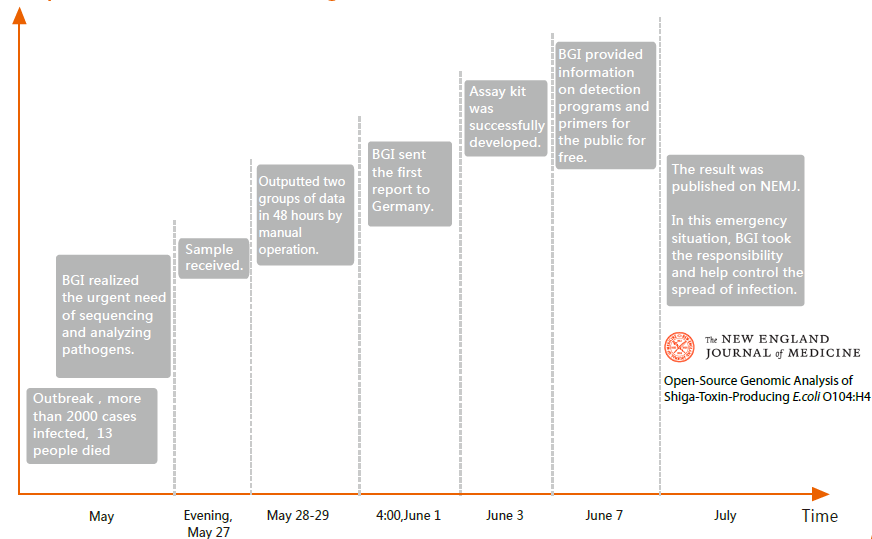
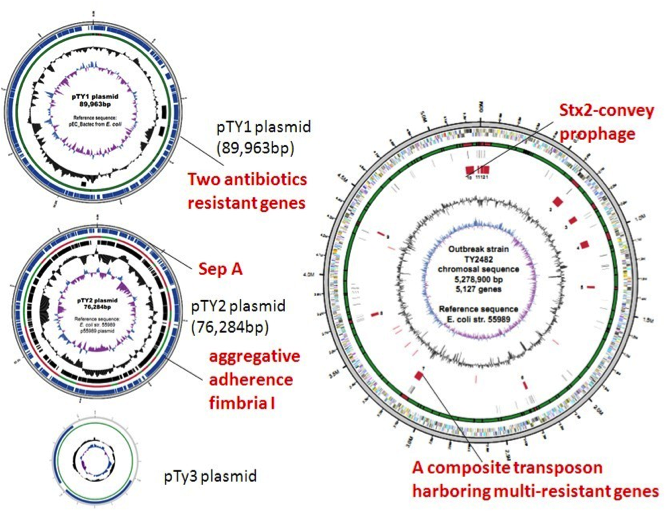
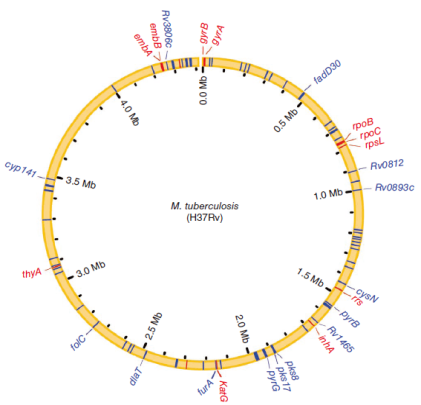
Figure 4. Genomic locations of the 85 drug resistance–associated genes.Genes previously reported to be associated with drug resistance are shown by red lines, whereas those associated with drug resistance for the first time in this study are shown by blue lines.
Industrial:
 Microorganisms are applied in a wide range of applications in industry, such as dairy processing, fermentation and bioreactors. Genetic studies on microorganisms can assist in identifying bioactive substances related pathways, bioenergy pathways and directional selection of microorganisms in different industrial conditions. BGI applies meta-omics research to help customers understand the diversity of microbial species and screen the functional microbes in food fermentation, such as wine, soy sauce, pickled vegetable and so on.
Microorganisms are applied in a wide range of applications in industry, such as dairy processing, fermentation and bioreactors. Genetic studies on microorganisms can assist in identifying bioactive substances related pathways, bioenergy pathways and directional selection of microorganisms in different industrial conditions. BGI applies meta-omics research to help customers understand the diversity of microbial species and screen the functional microbes in food fermentation, such as wine, soy sauce, pickled vegetable and so on.
Agricultural:
Applying microorganisms in various agricultural fields has been reported in numerous studies. For example, Rhizobia are known to produce nitrogen for plants like legumes, and Bacillus thuringiensis is used as a biological pesticide because of the Bt toxin produced by the bacteria. Furthermore, Genomics studies provide essential information on understanding crop pathogens as well as beneficial microbes. For instance, Stripe rust, caused by Puccinia striiformis f. sp. tritici (Pst), is one of the most destructive diseases of wheat. Recently, we generated a significantly improved draft genome (~110 Mb) of a Chinese Pst isolate, CY32, using a ‘fosmid-to-fosmid’ sequencing strategy that combines the use of a fosmid library and Illumina sequencing. In addition, we resequenced five prevalent Pst isolates collected from different geographical locations. We identified a large number of SNPs using whole-genome comparative analyses between Pst-CY32 and individual re-sequenced isolates (Figure 5, paper).
Figure 5. Phylogenic tree of 7 Pst isolates constructed with their cSNPs using Mega5-nj. The origin and isolation date of each isolate were marked.
Environmental:
 Microorganisms play a potential significant role in addressing environmental issues, energy supply and natural resource conservation. In an effort to systematically determine the functional and evolutionary diversity of microbial communities across the globe, BGI has assumed a leading role in the Earth Microbiome Project, an international research collaboration to analyze 200,000 samples using metagenomics to produce a global Gene Atlas. BGI metagenomics services allows for sequencing of the genomic DNA directly from microbial communities, bypassing microbe isolation and cloning steps.
Microorganisms play a potential significant role in addressing environmental issues, energy supply and natural resource conservation. In an effort to systematically determine the functional and evolutionary diversity of microbial communities across the globe, BGI has assumed a leading role in the Earth Microbiome Project, an international research collaboration to analyze 200,000 samples using metagenomics to produce a global Gene Atlas. BGI metagenomics services allows for sequencing of the genomic DNA directly from microbial communities, bypassing microbe isolation and cloning steps.

