Technology Platforms

BGI uses state-of-the-art technologies from numerous leading manufacturers and suppliers to provide reliable genomic, transcriptomic, epigenomic, metagenomic and proteomic services. Powered by over 230 sequencers, mostly on Illumina HiSeq, Complete Genomics and Ion Torrent platforms, with the data output capability of ~16T/day, we provide industry-leading sequencing capacity and turnaround time. BGI also provides extensive bioinformatics analyses supported by several supercomputing centers to convert data into knowledge. All of BGI’s services offered on the following technology platforms are carefully monitored by strict internal quality control and data security systems.
- Complete Genomics Platform
- Ion Proton Platform
- Illumina HiSeq 2000
- Bionano Irys System
- Roche GS FLX Platform
- Ion Torrent
- Applied Biosystems 3730xl DNA Analyzer
Proteomics Mass Spectrometry Platform:
- Thermo Scientific LTQ Orbitrap Velos
- Bruker maXis
- AB Sciex QTRAP 5500
- Bruker Ultrafle Xtreme
- Waters Xevo TQ-S
Sequencing Platform
Complete Genomics Platform
Complete Genomics platform provides industry-leading sequencing quality, and comprehensive analysis of all variation types, with two advanced proprietary sequencing technologies: DNA nanoball arrays (DNB arrays) and combinatorial probe-anchor ligation (cPAL). It supports a wide range of applications such as whole human genome sequencing, whole exome sequencing, and clinical diagnostics.
Highlights:
- Industry-leading accuracy reads (1 error in 100,000 bases) enabled by proprietary library construction in conjunction with combinatorial probe-anchor ligation technology
- Massively parallel DNA nano-arrays to minimize reagent usage and maximize imaging efficiency
- Sensitive detection of allele heterozygosity with proprietary local de novo variant mapping and assembly software
- Comprehensive results spanning all variation types including: SNPs, indels, CNVs, SVs, and mobile element insertions
- High specialized pipeline and informatics for human genome research with over 15,000 genomes analyzed so far
Ion Proton Platform
The proton uses semiconductor technology and real time measurement of hydrogen ions to translate chemical information directly into digital form, eliminating the need for expensive optics and complex sequencing chemistries. This instrument provides flexible sequencing options for generating up to 12-13 Gb of data per run with sample multiplexing. The Proton is ideal for exome sequencing, transcriptome sequencing, amplicon re-sequencing, targeted capture re-sequencing, microbial and viral whole genome sequencing.
- Fastest turnaround time with only 2-4 hours of sequencing run time
- Proprietary semiconductor sequencing technology with biochemistry to directly translate chemical information into digital data
- Scalable and flexible throughput of 1-2 human exomes or 1-6 RNA-Seq (Quantification) samples per sequencing run
- Over 150bp of average reads length on majority of reads
Illumina HiSeq 2000
The Illumina HiSeq 2000 system is a rapid, high-throughput, next-generation sequencing platform that supports the following applications: de novo sequencing, whole genome resequencing, RNA sequencing, disease research, epigenomics, microbial sequencing, and others.
Highlights:
- Highest sequencing output up to 55 Gb per day for 2 x 100 bp run and up to six billion paired-end reads per run.
- Parallel sequencing of millions of fragments using proprietary chemistry
- Four-color DNA sequencing-by-synthesis technology
- High accuracy and true base-by-base sequencing, providing the greatest yield of perfect reads and bases greater than Q30
- Lower cost due to bypassing cloning, transformation, and plasmid preparation
Bionano Irys System
The fully automated Irys benchtop instrument uses the IrysChip to uncoil and confine long DNA molecules in Nanochannel Arrays™ where they are uniformly linearized in a highly parallel display for high-resolution, single-molecule imaging. The result is sequence information over extremely long “reads” ranging from hundreds of kilobases to a megabase, where the sample’s valuable structural information is preserved. The system is ideal for de novo assembly of genomes containing complex, highly variable regions, as well as the accurate detection of all types of structural variation, both balanced and imbalanced, within complex heterogeneous samples.
Highlights:
- Fully automated cycling of DNA into chip and imaging
- 3 laser lines for simultaneous labeling of multiple events
- High-resolution optics with proprietary Auto-Focus system
- 50-100 GB/chip throughput
- Automatic de novo assembly and structural variation detection
Roche GS FLX Platform
The GS FLX+ System features the unique combination of long reads, exceptional accuracy and high-throughput, making the system well suited for larger genomic projects. Advancements in sequencing chemistry, instrumentation and software offer the latest improvements in GS FLX System performance, with read lengths up to 1 kb. Available as an on-site instrument upgrade or new instrument, the GS FLX+ System is designed for use with both the new long-read Sequencing Kit XL+ and existing Sequencing Kit XLR70.”
Ion Torrent
Ion Torrent integrates fluidics, micromachining, and semiconductor technology, and provides rapid DNA sequencing with quality data. Ion Torrent technology identifies DNA sequence by measuring the change of PH value during DNA synthesis. Ion Torrent currently offers three types of chips which generate various data throughput without two hours. BGI recently used the Ion Torrent to rapidly sequence the deadly strain of the German E coli outbreak.
Applied Biosystems 3730xl DNA Analyzer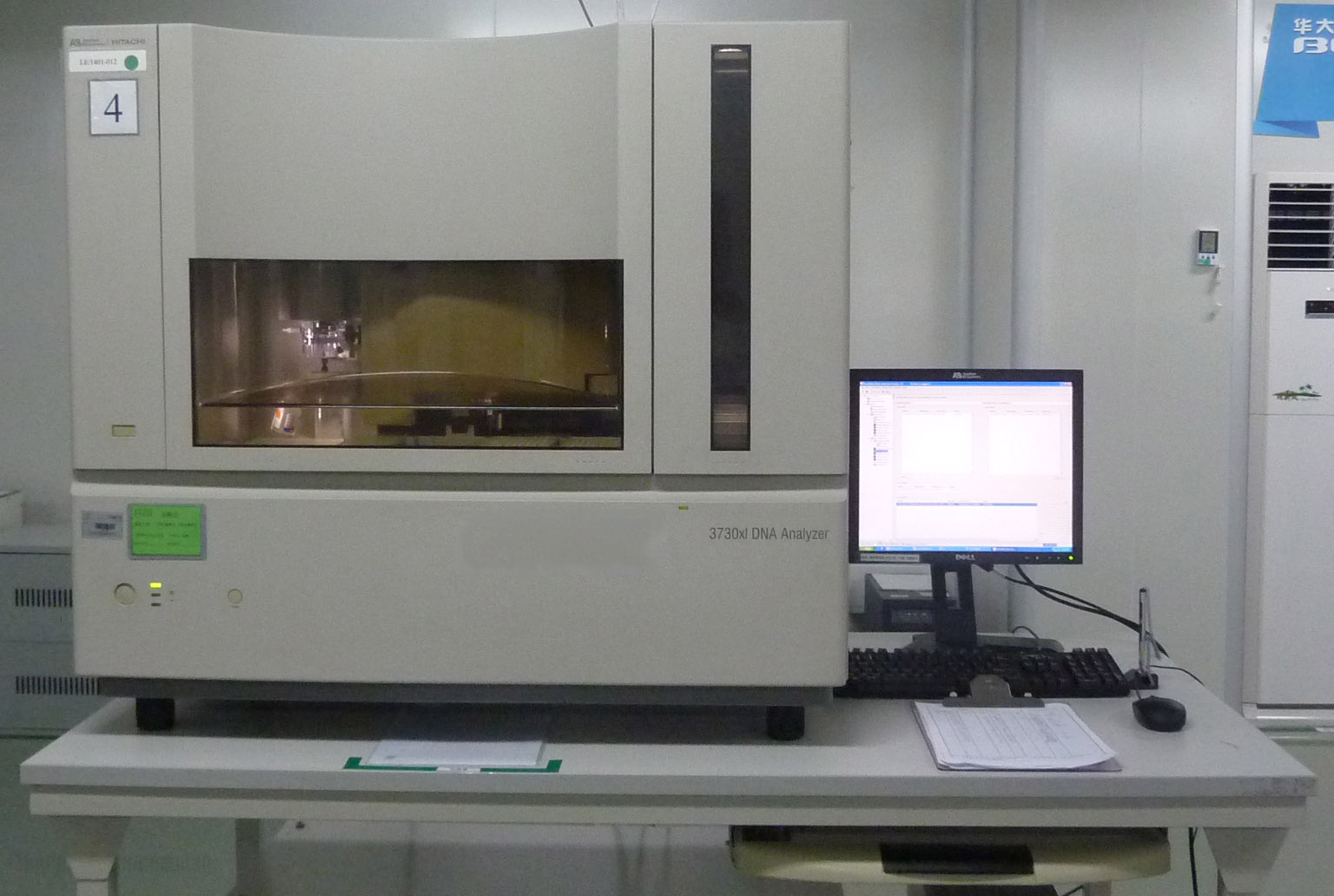
The 96-capillary 3730xl DNA Analyzer uses the latest optical design, a more optimized reagent mix, and fully automated operational processes to improve productivity and data quality while reducing the maximum consumption of reagents. Features include:
- Ease of use
- Built-in integrated autosampler and sample hole punch device
- Internal automated bar code identification
- High-throughput results
- Capacity to run up to 9,600 samples on one kit
- High-quality sequencing results
- Electrophoresis temperature of 70°C to remove the impact of secondary structure
- A new gel for measuring the length of 1,100 sequenced base pairs
- Base identification and quality score software to improve sequencing accuracy
- High-sensitivity optical system
- Capillary fluorescence detection
- Post-thin CCD detection system
- Double-beam laser excitation of bilateral, highly homogeneous fluorescence signal intensity
- High-sensitivity of detection of DNA in the concentration range
Exome Capture Platform
Agilent Sure Select Target Enrichment
Agilent SureSelect platform enables next-gen sequencing on key genomic regions of interest while reducing cost per sample. Agilent’s SureSelect platform currently supports target enrichment on the Illumina GA and HiSeq, Applied Biosystems SOLiD, and 454 Life Sciences FLX and GS Junior NGS systems. Protocols and kits have been optimized for in-solution work that are custom designed, along with a catalog product targeting the human exome. The platform is easily scalable for large, high-throughput studies and introduces little to no bias.
Roche NimbleGen Sequence Capture Technology
NimbleGen Sequence Capture technology is a revolutionary single step process for the enrichment of selected genomic regions from full complexity human genomic DNA. Sequence Capture was developed to eliminate the necessity of setting up thousands of PCR reactions, instead allowing for parallel enrichment of target regions in a single experiment. In combination with high throughput sequencing (short- or long-read), Sequence Capture has made targeted sequencing possible and accessible to more life science researchers. Two options are available for targeted enrichment of genomic regions:
- SeqCap EZ Library is a solution-based capture method that enables enrichment of the whole exome or customer regions of interest in a single test tube.
- Sequence Capture Arrays enable researchers to target custom regions of interest in the human genome.
Genotyping Platform
 Illumina iScan
Illumina iScan
At the System’s core is the iScan Reader, which incorporates high-performance lasers, optics, and detection systems. The iScan Reader offers sub-micron resolution and unmatched throughput rates. Even the highest density BeadChips can be scanned in just minutes, allowing processing of up to 96 multi-sample BeadChips per day.
- These imaging features are especially beneficial for high-density genotyping, CNV analysis, DNA methylation, and gene expression profiling
- The iScan System supports Infinium, GoldenGate, DASL, Gene Expression (Direct Hybridization), and Methylation assays along with different multi-sample BeadChip formats
Sequenom MassARRAY iPLEX® Gold
iPLEX® Gold is a leading technology for SNP Genotyping. The MassARRAY system is widely used for fine mapping and validation of GWAS studies linkage studies, as well as routine genetic testing of SNP panels of interest. MassARRAY SNP Genotyping combines the benefits of a simple and accurate primer extension chemistry with state of the art MALDI-TOF mass spectrometry to quickly and cost effectively characterize genotypes with the highest levels of reproducibility. The iPLEX Gold assay used in hundreds of publications to routinely design assays at a multiplexing level of up to 40-plex, offering a high level of flexibility.
Optical Mapping Platform
OpGen Argus Optical Mapping System
De Novo Technology — Generate Microbial Whole Genome Maps from Single DNA Molecules
- a non gel-based, PCR-based, or sequencing based methodology
Complete System — Includes everything needed to make an Optical Map
- hardware, software and consumables
Software for analyzing entire Microbial Genomes
- strain typing, comparative genomics, whole genome sequence assembly
Proteomics Mass Spectrometry Platform
Thermo Scientific LTQ Orbitrap Velos
The LTQ Orbitrap Velos system offers ultra-high resolution and accurate mass data for precise intact protein analysis. More unique proteins can be identified in a single experiment using this instrument, while delivering optimal structural characterization and rapid quantitation of low level isobarically-labeled peptides. The LTQ Orbitrap Velos has high mass accuracy that increases the speed and confidence of identification in complex samples. This mass spectrometry supports a mix of applications, including proteomics, drug discovery and development, and metabolomics.
Bruker maXis
MaXis is a novel high-speed, ultra high resolution mass spectrometer, with an acquisition speed of 20 Hz full spectra per second. It has a variety of analytical applications such as small molecule identification, metabolomics, quantitative proteomics, biomarker discovery and validation. With unparalleled sensitivity and speed, the maXis offers maximum flexibility from small molecule to intact large molecules in both MS and MS/MS modes.
AB Sciex QTRAP 5500
 QTRAP 5500 is a highly-sensitivity triple quadrupole system that allows multiple reaction monitoring to be performed. It offers quicker and easier identification, characterization, and quantitation of metabolites. QTRAP 5500 has applications in drug discovery, ADME/Toxicology research, metabolomics, forensic, clinical research, environment and food security analysis. This next-generation technology is equipped with a scan speed of up to 2400 amu/s (QQQ) and up to 20,000 amu/sec (Trap), generating unparalleled results.
QTRAP 5500 is a highly-sensitivity triple quadrupole system that allows multiple reaction monitoring to be performed. It offers quicker and easier identification, characterization, and quantitation of metabolites. QTRAP 5500 has applications in drug discovery, ADME/Toxicology research, metabolomics, forensic, clinical research, environment and food security analysis. This next-generation technology is equipped with a scan speed of up to 2400 amu/s (QQQ) and up to 20,000 amu/sec (Trap), generating unparalleled results.
Bruker Ultrafle Xtreme
Ultrafle Xtreme offers high throughput analysis and a top-down protein sequencing method without enzymatic digestion. This MALDI-TOF/TOF mass spectrometer is the only instrument that has a combination of full 1 kHz smartbeam II laser in both TOF and TOF/TOF modes with ultra-high performance and extreme flexibility. Ultrafle Xtreme supports the following applications: 2DE spots identification, LC-MALDI proteome profiling, LC-MALDI iTRAQ quantification, unknown protein de novo analysis, and MALDI-Imaging.
Waters Xevo TQ-S
 The Xevo TQ-S tandem mass spectrometer is able to accurately measure target compounds in samples at femtogram levels or below. This system features a new StepWave ion-transfer technology, Scanware, and RADAR . These combined features deliver class leading UPLC/MS/MS sensitivity, provide rapid, high quality, UPLC compatible, MS/MS acquisition, and allow collection of highly specific quantitative data for target compound, respectively. Xevo TQ-S has applications such as quantitative analysis, food safety, environmental monitoring, and forensic toxicology.
The Xevo TQ-S tandem mass spectrometer is able to accurately measure target compounds in samples at femtogram levels or below. This system features a new StepWave ion-transfer technology, Scanware, and RADAR . These combined features deliver class leading UPLC/MS/MS sensitivity, provide rapid, high quality, UPLC compatible, MS/MS acquisition, and allow collection of highly specific quantitative data for target compound, respectively. Xevo TQ-S has applications such as quantitative analysis, food safety, environmental monitoring, and forensic toxicology.