
de novo Sequencing
Cases
Technical Information
Contact Us
 de novo sequencing provides the first genome sequence of an organism. With the advent of rapid, low-cost next-generation sequencing technology researchers can now obtain whole genome data for organisms previously considered too low a priority to sequence. The availability of this whole genome data has allowed large-scale genomic studies to be performed that were unimaginable just a few years ago. BGI and its global collaborators have now initiated 505 plant and animal genome projects, completed fine or draft genome maps for nearly 100 species and finished the sequencing of about 200 species. The completed projects include rice, silkworm, cucumber, panda, camel, oyster, ant genomes, and more.
de novo sequencing provides the first genome sequence of an organism. With the advent of rapid, low-cost next-generation sequencing technology researchers can now obtain whole genome data for organisms previously considered too low a priority to sequence. The availability of this whole genome data has allowed large-scale genomic studies to be performed that were unimaginable just a few years ago. BGI and its global collaborators have now initiated 505 plant and animal genome projects, completed fine or draft genome maps for nearly 100 species and finished the sequencing of about 200 species. The completed projects include rice, silkworm, cucumber, panda, camel, oyster, ant genomes, and more.
Benefits:
- More comprehensive maps of genetic variation
- Variable gradient insert libraries enable fine mapping of the genome
- Reliable genome assembly by BGI’s independently developed software- SOAPdenovo
- NGS high-throughput sequencing reduces cost
- Experienced bioinformatics team
The Sequence and de novo Assembly of the Giant Panda Genome. Nature 2010.463:311-317.

The panda genome was the first genome completely sequenced by using next generation sequencing platform alone. It provides clues to the understanding of everything from the panda’s strict bamboo diet to it’s genetic diversity. It may also aid in the panda conservation in the future.
Genome Sequence and Analysis of the Tuber Crop Potato. Nature. 14 Jul 2011; 475:189-195.

Potato (Solanum tuberosum L.) is the world’s most important non-grain food crop and is central to global food security. It is clonally propagated, highly heterozygous, autotetraploid, and suffers acute inbreeding depression. Here we use a homozygous doubled-monoploid potato clone to sequence and assemble 86% of the 844-megabase genome. We predict 39,031 protein-coding genes and present evidence for at least two genome duplication events indicative of a palaeopolyploid origin.
The Genomic Sequence of the Chinese Hamster Ovary (CHO)-K1 Cell Line. Nature Biotechnology. 31 Jul 2011; 29: 735–741.

Chinese hamster ovary (CHO) cell lines are the preferred and most commonly used mammalian cell line today in biological and medical research. With the advanced genome sequencing capability of BGI, the CHO-K1 ancestral cell line was sequenced by de novo sequencing and then assembled by BGI's Short Oligonucleotide Analysis Package, SOAPdenovo, resulting in the assembly of 2.45 Gb of the genomic sequence. This information was combined with transcriptome sequence data and led to the generation of 24,383 predicted genes. The study yields a better understanding of the genetics of CHO cells and will accelerate the discovery and development of new recombinant protein therapies.
Structural Variation in Two Human Genomes Mapped at Single-Nucleotide Resolution by Whole Genome de novo Assembly. Nature Biotechnology. 2011.29:723-730.
Whole genome de novo assembly was used to map structural variations in an Asian and an African genome. Small-and-intermediate-size homozygous variants including insertions, deletions, and inversions were identified. These findings also demonstrate that whole genome de novo sequencing is a practical approach to deriving more comprehensive maps of genetic variation.
Workflow:
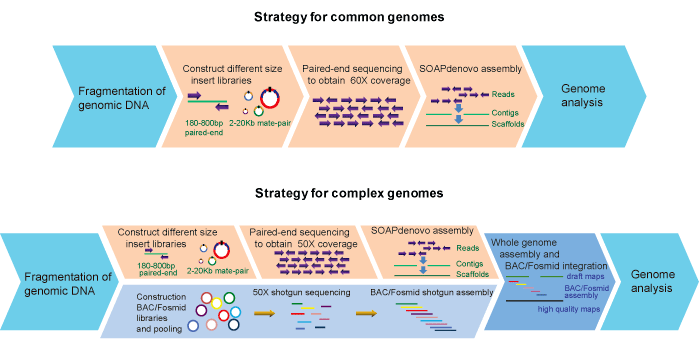
Bioinformatics:
Genome analysis: Genome size, GC content, heterozygous rate, repeats content, sequence depth, autosomal and gene region coverage evaluation.
Genome annotation: Repeat, ncRNA annotation, gene prediction, gene function annotation.
Comparative genomics and evolution analysis: Orthologous gene clusters, phylogenetic analysis, divergence time and substitution rate estimation, whole genome alignment, segmental duplication, and conserved element.
Microbial de novo Sequencing:
For the genomic DNA samples you provide us:
- Purity: OD260/280=1.8-2.0
- Concentration: ≥50 ng/μl
- DNA amount: single library preparation starts from at least 6ug; 2 Kb~6 Kb
Large-insert library preparation starts from 40ug; and PCR-free library preparation (for the genome with rather high or low GC content) starts from 30ug.
The total amount should be determined case by case.
Plants and Animals de novo Sequencing:
For the genomic DNA samples you provide us:
- Purity: OD260/280=1.8~2.0, without protein, RNA, or other visible contamination
- Concentration: for short-insert libraries, ≥50 ng/μl. For 2 Kb~10 Kb large-insert libraries, ≥150 ng/μl; for 20 Kb large-insert libraries, ≥200 ng/μl;
- DNA amount: for short-insert libraries, single library preparation starts from 6 μg; for 2 Kb~10 Kb large-insert libraries, ≥40 μg; for 20 Kb large-insert libraries, ≥60 μg; the total sample amount for whole genome sequencing is about 500 μg~1 mg;
- Sample quality: genomic DNA should be intact. If large-insert libraries (>5 Kb) should be constructed, genomic DNA fragments distribution should be 23 Kb or above according to electrophoresis gel result. For the result of pulsed-field gel electrophoresis, the main bands of DNA should be more than 40 Kb.

