
Bioinformatics Software
BGI has developed a series of bioinformatics analysis tools for various applications. SOAP (Short Oligonucleotide Alignment Program) has been in evolution from a single alignment tool to a package that provides full solution to next generation sequencing data analysis, and has been widely adopted by more than 10,000 users. BGI also applies a variety of open source software, for example ABySS and Velvet, in order to provide comprehensive bioinformatics analysis for our sequencing services.
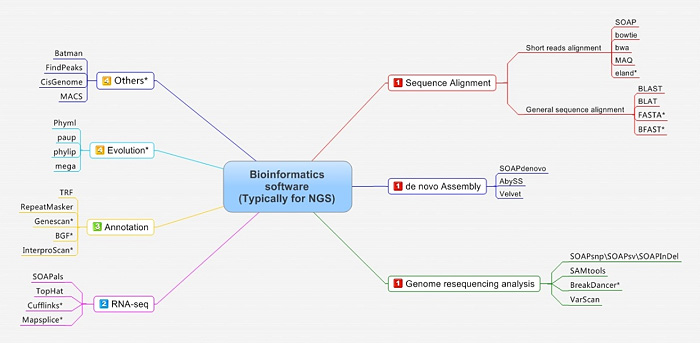
Figure 2. Bioinformatics software (typically for NGS) employed by BGI.
The following is a list of software developed by BGI:
SOAPdenovo - SOAPdenovo, a short read de novo assembly tool, is a package for assembling short oligonucleotide into contigs and scaffolds. SOAP family software can be found here (http://soap.genomics.org.cn/).
RePS (repeat-masked Phrap with scaffolding) - RePS is a WGS sequence assembler. It identifies repeated kmer sequences and deletes WGS sequence prior to assembly. The established software Phrap is used to compute meaningful error probabilities for each base. Clone-end-pairing information is used to construct scaffolds that order and orient the contigs. The updated version of RePS incorporates some of the ideas introduced by Phusion on clustering.
Exon_Capture_Pipeline - Whole-genome exon trapping analysis software.
Maq (Mapping and Assembly with Quality) - Maq builds assemblies by mapping short reads to reference sequences. Maq was previously known as mapass2.
ReAS - Software to recover ancestral sequences for transposable elements using unassembled reads from a whole genome shotgun sequencing.
SOAPaligner/soap2 - SOAPaligner/soap2 is a program for faster and more efficient alignment for short oligonucleotide onto reference sequences. SOAPaligner/soap2 is compatible with numerous applications, including single-read or pair-end resequencing.
SOAPsnp - SOAPsnp is an accurate consensus sequence builder based on Soap1 and SOAPaligner/soap2′s alignment output. It calculates a quality score for each consensus base, which can be used for any latter process to call SNPs.
SOAPindel - SOAPindel is developed to find the insertion and deletion specially for re-sequence technology.
SOAPsv - SOAPsv is a program for detecting the structural variation.
SOAP3/GPU - SOAP3 is a GPU-based software for aligning short reads with a reference sequence. It can find all alignments with k mismatches, where k is chosen from 0 to 3. When compared with its previous version SOAP2, SOAP3 can be up to tens of times faster.
MIEREAP - This is used to identify both known and novel microRNAs from small RNA libraries that were deeply sequenced using Illumina-Solexa/454/Solid technology.
FGF - (Fishing Gene Family, http://fgf.genomics.org.cn/) – This finds gene families, plots phylogenetic trees, and provides evolutionary information to gene duplication.
SVBP - This provides reliability tests and results visualization for sequence assembly.
WEGO - (Web Gene Ontology Annotation Plot, http://wego.genomics.org.cn/cgi-bin/wego/index.pl) – Web Gene Ontology Annotation Plot is a useful tool for plotting GO annotation results especially for comparative genomics.
HIBAIS - Ancestor deduction software based on HapMap.
SOLEXA-MRNATAG_PIPELINE - Digital gene expression software based on Illumina-Solexa sequencing data
CAT (Cross-species Alignment Tool) - Allows mRNA sequence and mammalian genome alignment across species
KaKs_Calculator - This calculates nonsynonymous (Ka) and synonymous (Ks) substitution rates. More information is available here http://evolution.genomics.org.cn/software.htm.

