
- Genomics
- Transcriptomics
- Epigenomics
- Meta-omics
- Proteomics
- Single-Cell Sequencing
- Immune Repertoire Sequencing
- FFPE Samples

Single Sample Sequencing
BGI provides sequencing services for bacteria (bacteria, fungi, bacteria liquid puncture flat), plasmid, and PCR products (the original and purified). We also provide services such as plasmid extraction, purification of the PCR product, designing the sequencing primers, DNA sequence data analysis, long fragment walking, sequence assembly, etc.
Workflow:
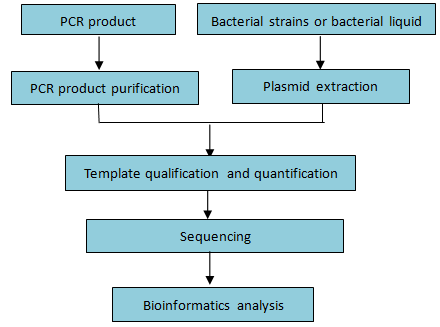 Figure 8. Workflow of single sample sequencing
Figure 8. Workflow of single sample sequencingBioinformatics Contents:
Bioinformatics contents consist of basic data analysis processing, including base calling and the sequencing results of classification, etc.
Sample Requirements:
Bacterium fluid
-
Sample amount for fresh bacterium fluid: 500-1000 μL or 1-2mL precipitation bacteria that can extract plasmid directly
PCR Liquid (Unpurified)
- Sample amount: ≥ 25 μL
- Sample concentration: ≥ 20 ng/μL
Plasmid
- Sample amount: volume ≥ 20 μL
- Sample concentration: ≥ 100 ng/μL
Turnaround Time:
The turnaround time for plasmid and PCR results using the standard procedure (above) is about 24 hours. The turnaround time for bacterium fluid results using the standard procedure (above) is about 48 hours.
Completion Milestone:
Credible read lengths can reach 800bp (except natural termination), and the results of the first round of sequencing reaction can be completed within three working days.
Frequently Asked Question:
Q1: What is the basis of DNA sequencing? Describe the accuracy of sequencing using the ABI3730xl sequencer?
A: The Sanger sequencing method is the basis of DNA sequencing. The specifics of the method can be found in “Molecular Cloning.”
The ABI3730 sequencer uses AB sequencing reagents and interprets bases sequencing using its own judgment value (Quality Value). Depending on the size of the QV value, it is possible to determine the accuracy of each base. ABI’s commitment to the sequencing length of the standard sample is 929bp, and the data correlation to the Q20 standard is more than 80%.
Q2: How can we avoid bacteria growth when shipping samples?
A: Problems with bacteria can be a result of the transportation time. If you are sending your samples over a long distance, we recommend that you send puncture bacteria. Also, some strains are unique and require special media and environmental training. If this is the case, we recommend that you send 2ml of fresh bacteria or plasmid.
We recommend DH5α and DH1 and C600Ecoli strains as the host bacteria. The XL1-Blue E.coli bacteria are also acceptable, but its growth is too slow. E.coli bacterial strains of the JM series, TG1 and HB101 series produce large amounts of carbohydrates. Using these strains should be avoided if possible.
Q3: Samples identified before being sent to your company for sequencing showed that the target fragments were inserted into the plasmids. However sequencing results showed no inserts in the plasmids? Why did this occur and will we be charged for these results?
A: These results can be caused by the following two reasons:
1) The identification showed a false positive result. The inserted fragment can be identified in two ways: PCR and enzyme digestion verification. PCR can be influenced by many factors resulting in more false positives than compared with enzyme digestion verification. However, if contamination or other problems occurred during the cloning step, it can show the same size band as a target fragment from the negative clone.
2) Target fragments can be lost during the culture process. Due to many limitations, we cannot culture samples under certain conditions, and this may result in the lost of target fragments. We cannot anticipate this situation.
We do charge for these results.
Q4: When I performed PCR, the PCR primers worked very well. Why are the sequencing results not as good when using the same primers?
A: It should be made clear that not all primers for PCR can be used for sequencing. These following kinds of primers are not suitable for sequencing.
1) Degenerate primers: There are several binding sites on the sequencing template for degenerate primers that will directly influence sequencing results.
2) Random primers: The lengths of random primers (e.g. RAPD primers) are often very short and the annealing temperature is low. As a result, it cannot bind to the template very well during the sequencing reaction.
3) Overlong primers: Normal sequencing primers are less than 24bp. (The longest is 30bp.) Overlong primers will bind to the multi-locus on the template resulting in an increase in the background for sequencing. Furthermore, the purity of longer primers cannot be guaranteed. The purity of primers for sequencing is normally larger than 90% which will significantly increase the background of sequencing reaction when the purity is low. This directly influences the sequencing result.
4) Labeled primers: Here we are referring to fluorescent labeled primers. The four types of bases we use for the sequencing reaction are labeled with fluorescence, and the labeled primers will affect the results. Other larger gene labeled primers are also not suggested for sequencing as this labeled group will directly affect the migration rate of DNA fragments, resulting in incorrect peak shapes for sequencing.
5) Impure primers: Purity is extremely important for sequencing primers as non-full length fragments in synthesis primers can result in a strong background. As an example, given 20bp sequencing primers, the purity will be 70% at most if it is purified by directly de-salting. This means that 30% of the primers will be measured as background noise, greatly influencing the sequencing result. Generally, primers that are purified using PAGE or OPC will meet the needs for sequencing.
Q5: How do you deal with the sequencing samples and primers after finishing the sequencing?
A:If you want the sequencing samples and your primers returned, please indicate that in your order, and we will return them when the order is complete. We store the samples and primers for one month from the date received. As such, if you want to sequence samples that were sent more than a month ago, you will need to supply the samples again.
Q6: What causes a weak signal or no signal in sequencing?
A: There are four main reasons that can affect the signal.
1) The template quality is poor.
2) Primer and template sequences don’t match.
3) Primer parameters do not comply with the standard of sequencing primers.
4) There is a secondary structure in the sequence.
Frequently during our sequencing we encounter a number of failures due to a lack of template primer binding sites. Some of the reasons for this include:
1) The customer provided incorrect information for the vector or the primers.
2) The PCR primer was used for sequencing the plasmid, but the plasmid was an empty carrier.
If the carrier is transformed or the template has mutations, it can lead to a loss of the original primer binding sites.
