
- Genomics
- Transcriptomics
- Epigenomics
- Meta-omics
- Proteomics
- Single-Cell Sequencing
- Immune Repertoire Sequencing
- FFPE Samples

PCR Re-sequencing
Target sequences are amplified in different individuals with PCR. Then the amplified products are sequenced and analyzed. PCR re-sequencing has the following applications:
- Single-gene genetic diseases: pathogenic gene research and gene diagnosis;
- Complex diseases: relevant gene research and drug susceptibility analysis;
- Association analysis between the phenotypes and nucleic acid level variation with mutation, insert, delete, etc.
Workflow:
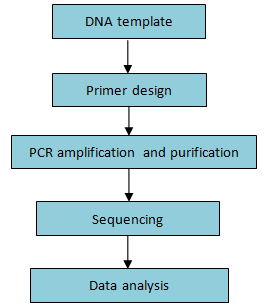 Figure 5. Workflow of PCR re-sequencing
Figure 5. Workflow of PCR re-sequencingBioinformatics Content:
Bioinformatics contents consist of basic analysis based on the PloyPred software.
Sample Requirements:
- DNA templates and primers required for PCR or sequencing:
Requirements are a genome DNA concentration of 0.1-1 μg/µL, with a volume of ≥ 20 μL per PCR reaction. The primers should be dry powder (1 OD).
- PCR products:
Sample concentration should be ≥ 50 μg/µL for unpurified PCR product and ≥ 20 μL; for purified PCR product. The corresponding gel electrophoresis result should be provided.
Turnaround Time:
The turnaround time is determined by the number of samples and sequences.
Completion Milestone and Result Submission:
Completion milestone:
Project completion is based on a quality score of Q 20.
Results submission:
Results consist of a sequencing peak figure, basic analysis, and the final report.
